CAGI2 Challenge
The second round of CAGI was completed in 2011, with 11 challenges provided to predictors. The experiment yielded a total of 117 predictions from 21 groups from 18 countries, with 55 people attending the December 2011 meeting. Experimental results for the following CAGI 2011 challenges have been published: Mouse exomes challenge: Fairfield et al., Mutation discovery in mice by whole exome sequencing. Genome Biology 2011, 12:R86. doi:10.1186/gb-2011-12-9-r86
Cancer cell lines challenge: Heiser LM et al., Subtype and pathway specific responses to anticancer compounds in breast cancer. doi: 10.1073/pnas.1018854108. CBS challenge: Mayfield JA, et al. Surrogate genetics and metabolic profiling for characterization of human disease alleles. Genetics. 2012, 190(4):1309-23. doi:10.1534/genetics.111.137471. CAGI was presented at ISMB/EECB 2011 in a special session in July, covered in GenomeWeb and in a special session at the Pacific Symposium on Biocomputing in January 2012.
The flagship manuscript describing the meeting as well as a collection of papers describing the predictions and challenges is currently underway.
Key Dates
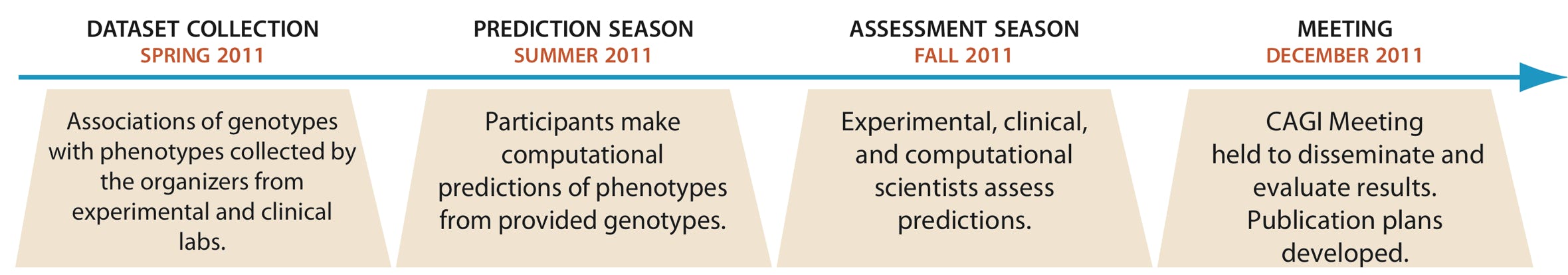
23 Jun 2011: Initial Datasets posted on the CAGI website
31 Aug - 11 Oct 2011: Prediction submission deadline depends on the dataset. The deadline is always 3PM EDT (Eastern time in the USA).
- 31 August - Mouse exomes
- 30 September - Cancer cell lines
- 3 October - CBS
- 4 October - MR-1, SCN5A
- 5 October - p53
- 6 October - Asthma
- 7 October - RAD50
- 21 October - riskSNPs
- 26 October - Crohn's disease
- 31 October - PGP
15 Oct 2011: Registration for CAGI 2011 Conference opens
9 - 10 December 2011: Two-day CAGI 2011 Conference at the University of California, San Francisco Mission Bay campus
Challenges
The following challenges were launched:
- Discordant monozygotic twins – identify the twin with asthma
- Breast cancer cell line pharmacogenomics dataset
- Cystathionine beta-Synthase (CBS) single amino acid mutations observed in patients
- Distinguish between exomes of Crohn's disease patients and healthy individuals
- Mouse exomes dataset – predict the causative mutation of a given phenotype
- The fitness of tagged transposon insertions in Shewanella oneidensis strain MR-1 under stress conditions
- Single amino-acid changes in the human p53 core domain that can restore activity of inactive p53 found in human cancers
- Personal genome project (PGP) - Predict individuals' phenotypes
- RAD50 variants in breast cancer patients and controls
- Novel NaV1.5 channel mutations associated with Brugada syndrome
- Identify the splicing impact of variation(Withdrawn)
- Molecular Mechanisms underlying associations between genetic variation and disease risk
Results
Personally identifiable data is restricted according to dataset requirements permanently. All datasets remain embargoed until published or otherwise released by providers. CAGI results are embargoed until 1 Jan 2012. CAGI 2011 Assessment results are available here for registered users.
Risk SNP Results
Personally identifiable data is restricted according to dataset requirements permanently. For results see here.
Organizers and Support
Organizers
Steven E. Brenner, CAGI Chair, University of California, Berkeley
John Moult, CAGI Chair, IBBR, University of Maryland
Susanna Repo, CAGI Organizer, University of California, Berkeley
Data Providers
Adam Arkin, University of California, Berkeley
George Church, Harvard Medical School
Andre Franke, Christian–Albrechts–Universität zu Kiel
Joe W. Gray, Lawrence Berkeley National Laboratory
Rick Lathrop and the p53 "cancer rescue" team, University of California, Irvine
John Moult and Lipika R. Pal, University of Maryland
Jasper Rine, University of California, Berkeley
Jeremy Sanford, University of California, Santa Cruz
Nicole Schmitt, University of Copenhagen
Jay Shendure, University of Washington
Michael Snyder, Stanford University
Sean Tavtigian, University of Utah
Confirmed Assessors
Rui Chen, Stanford University
Iddo Friedberg, Miami University
Gad Getz, Broad Institute
Sean Mooney, Buck Institute
Alexander A. Morgan, Stanford University
Artem Sokolov, University of California, Santa Cruz
Josh Stuart, University of California, Santa Cruz
Sean Tavtigian, University of Utah
Confirmed CAGI Advisory Board
Russ Altman, Stanford University
George Church, Harvard Medical School
Tim Hubbard, Wellcome Trust Sanger Institute
Scott Kahn, Illumina Inc.
Sean Mooney, Buck Institute
Pauline Ng, Genome Institute of Singapore
Confirmed CAGI Scientific Council
Patricia Babbitt, University of California, San Francisco
Atul Butte, Stanford University
Garry Cutting, Johns Hopkins University School of Medicine
Laura Elnitski, NIH/NHGRI
Reece Hart, Locus Development
Rachel Karchin, Johns Hopkins University
Robert Nussbaum, University of California, San Francisco
Michael Snyder, Stanford University
Shamil Sunyaev, Harvard Medical School
Joris Veltman, Radboud University Nijmegen
Liping Wei, Peking University
CAGI Assessors, ex officio
CAGI Advisory Board members, ex officio
Website Development and Administration
Maya Zuhl, IBBR, University of Maryland
Sri Jyothsna Yeleswarapu, Tata Consultancy Services
Support
The CAGI experiment website development has been supported in part by Tata Consultancy Services.
Funding for the CAGI 2011 Conference was made possible (in part) by 1R13HG006650-01 from the National Human Genome Research Institute. The views expressed in written conference materials or publications and by speakers and moderators do not necessarily reflect the official policies of the Department of Health and Human Services; nor does mention by trade names, commercial practices, or organizations imply endorsement by the U.S. Government.
Susanna Repo has been supported by Marie Curie International Outgoing Fellowship PIOF-QA-2009-237751.
CAGI 2011 Conference Slides and Videos
Slides and videos of the meeting presentations are available below. To view the videos, you need to have Quicktime installed on your computer. If you are using Windows, please go here to download the software: http://www.apple.com/quicktime/download/. On Macs, the software is already installed. Conference slides and videos are available only to logged in users. Please log in, in order to view the videos.
